Problem Statement
In Kruskal Algorithm, initially, all the nodes of the graph are separated from each other, which means they don’t have an edge between them. Then to obtain the minimum spanning tree from that graph we first sort the edges of the graph in a non-decreasing fashion. Then we pick the edges from left to right and connect the graph.
Now there are two possibilities, first one is if the current picked edge is already connected in the tree so in this case, we will just continue our process and if the current picked edge is not connected then we will just connect those two nodes with dsu (disjoint set union). In this manner, we finally conclude with the final MST of the given graph.
- Example:
- Input:
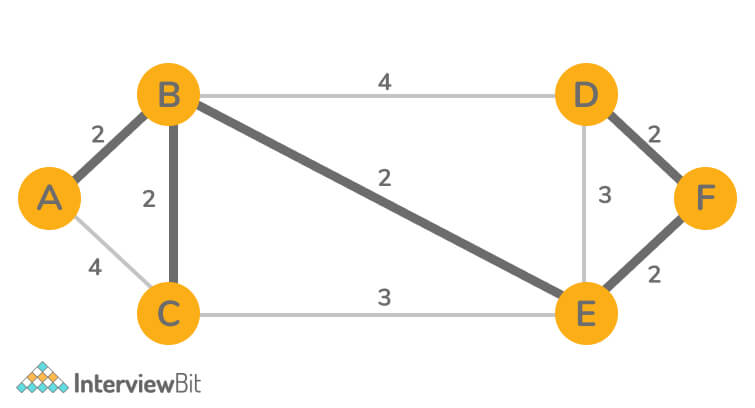
- Output: 10
Explanation: MST Weight = AB + BC + BE + BC+ EF = 10
Confused about your next job?
- Input:
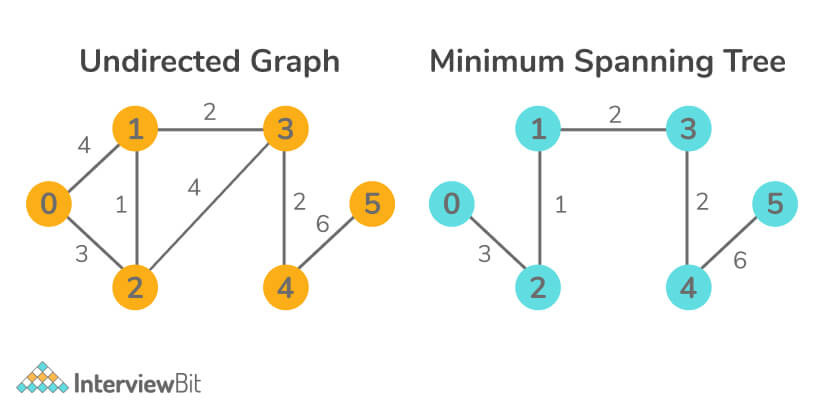
- Output: 14
Explanation: Here the second image is the MST image of the left image.
Approach
- Sort all the edges of the graph with respect to their weights. Initially sort the graph edges with respect to weights in a non-decreasing manner.
- Now traverse left to right means from smallest weight to largest weight and start adding edges to the MST.
- Add all those edges which don’t form a cycle, edges that connect only disconnected components.
So now one more question arises here, how will we check if the 2 vertices are connected or not?
To check if 2 vertices are connected or not we can use the dfs approach. In this, we first start our dfs from any 1st vertex and then check whether the 2nd vertex is visited or not. To run it efficiently we can do this by DSU (Disjoint Set Union).
Dry Run
Step 1:
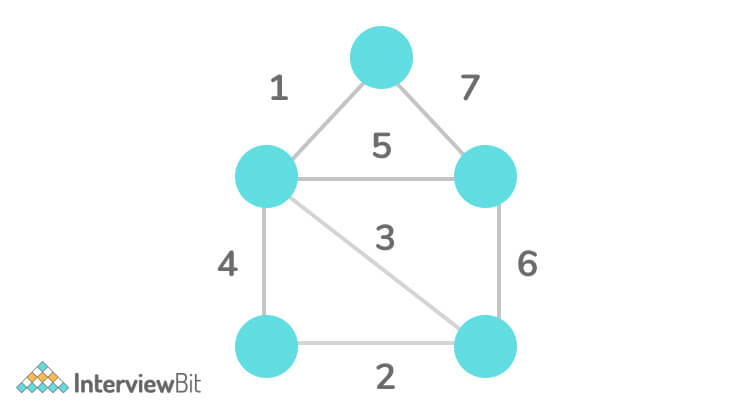
Step 2:
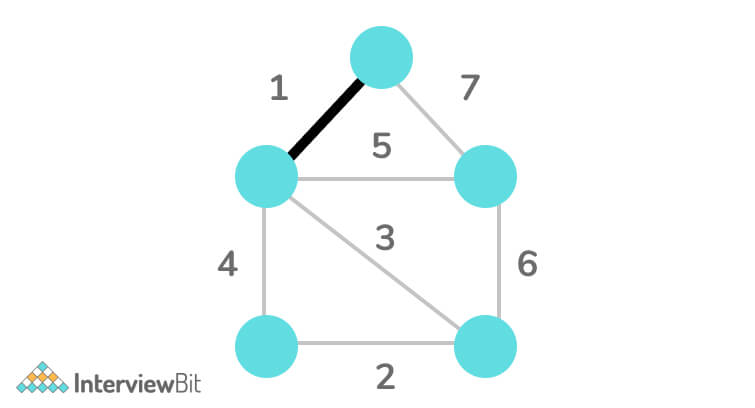
Step 3:
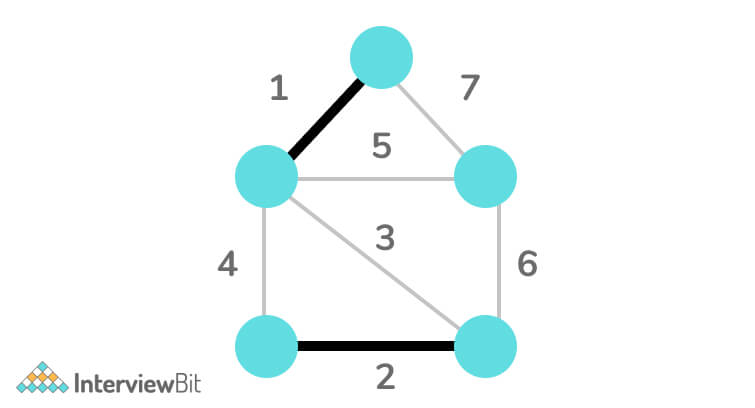
Step 4:
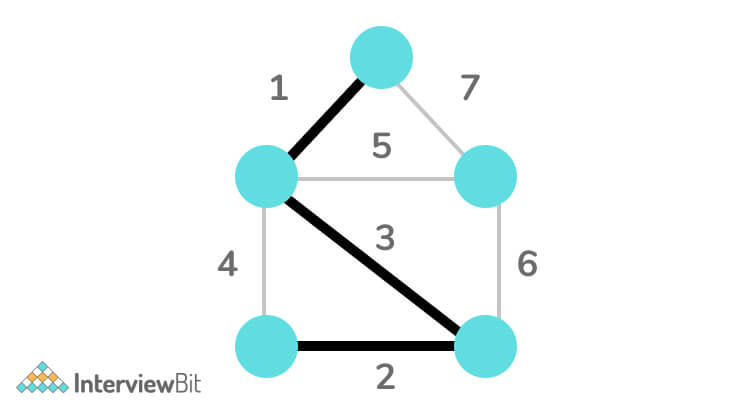
Step 5:
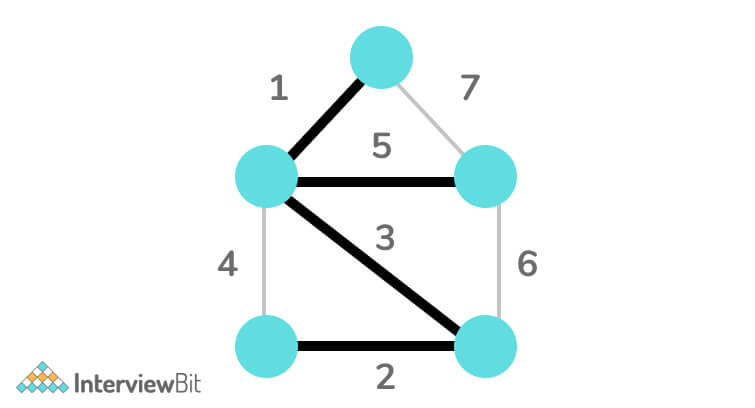
C/C++ Implementation
const int MAX = 1e4 + 5;
int id[MAX], nodes, edges;
pair <long long, pair<int, int> > p[MAX];
void initialize()
{
for(int i = 0;i < MAX;++i)
id[i] = i;
}
int root(int x)
{
while(id[x] != x)
{
id[x] = id[id[x]];
x = id[x];
}
return x;
}
void union1(int x, int y)
{
int p = root(x);
int q = root(y);
id[p] = id[q];
}
long long kruskal(pair<long long, pair<int, int> > p[])
{
int x, y;
long long cost, minimumCost = 0;
for(int i = 0;i < edges;++i)
{
// Selecting edges one by one in increasing order from the beginning
x = p[i].second.first;
y = p[i].second.second;
cost = p[i].first;
// Check if the selected edge is creating a cycle or not
if(root(x) != root(y))
{
minimumCost += cost;
union1(x, y);
}
}
return minimumCost;
}Java Implementation
class UnionFind{
final int[] parents;
int count;
public UnionFind(int n){
this.parents = new int[n];
reset();
}
public void reset(){
for(int i =0;i<parents.length;i++){
parents[i] = i;
}
count = parents.length;
}
public int find(int i){
int parent = parents[i];
if(parent == i){
return i;
}else{
int root = find(parent);
parents[i] = root;
return root;
}
}
public boolean union(int i, int j){
int r1 = find(i);
int r2 = find(j);
if(r1 != r2){
count--;
parents[r1] = r2;
return true;
}else{
return false;
}
}
}
class Solution {
public List<List<Integer>> findCriticalAndPseudoCriticalEdges(int n, int[][] edges) {
List<Integer>criticals = new ArrayList<>();
List<Integer> pseduos = new ArrayList<>();
Map<int[], Integer> map = new HashMap<>();
for(int i =0;i<edges.length;i++){
map.put(edges[i], i);
}
Arrays.sort(edges, (e1, e2)->Integer.compare(e1[2], e2[2]));
int minCost = buildMST(n, edges, null, null);
for(int i =0;i<edges.length;i++){
int[] edge = edges[i];
int index = map.get(edge);
int costWithout = buildMST(n, edges, null, edge);
if(costWithout > minCost){
criticals.add(index);
}else{
int costWith = buildMST(n, edges, edge, null);
if(costWith == minCost){
pseduos.add(index);
}
}
}
return Arrays.asList(criticals, pseduos);
}
private int buildMST(int n, int[][] edges, int[] pick, int[] skip){
UnionFind uf = new UnionFind(n);
int cost = 0;
if(pick != null){
uf.union(pick[0], pick[1]);
cost += pick[2];
}
for(int[] edge : edges){
if(edge != skip && uf.union(edge[0], edge[1])){
cost += edge[2];
}
}
return uf.count == 1? cost : Integer.MAX_VALUE;
}
}Python Implementation
class Solution:
def findCriticalAndPseudoCriticalEdges(self, n: int, edges: List[List[int]]) -> List[List[int]]:
def dfs(curr, level, parent):
levels[curr] = level
for child, i in graph[curr]:
if child == parent:
continue
elif levels[child] == -1:
levels[curr] = min(levels[curr], dfs(child, level + 1, curr))
else:
levels[curr] = min(levels[curr], levels[child])
if levels[child] >= level + 1 and i not in p_cri:
cri.add(i)
return levels[curr]
cri, p_cri = set(), set()
dic = collections.defaultdict(list)
for i, (u, v, w) in enumerate(edges):
dic[w].append([u, v, i])
union_set = UnionFindSet(n)
for w in sorted(dic):
seen = collections.defaultdict(set)
for u, v, i in dic[w]:
pu, pv = union_set.find(u), union_set.find(v)
if pu == pv:
continue
seen[min(pu, pv), max(pu, pv)].add(i)
w_edges, graph = [], collections.defaultdict(list)
for pu, pv in seen:
if len(seen[pu, pv]) > 1:
p_cri |= seen[pu, pv]
edge_idx = seen[pu, pv].pop()
graph[pu].append((pv, edge_idx))
graph[pv].append((pu, edge_idx))
w_edges.append((pu, pv, edge_idx))
union_set.union(pu, pv)
levels = [-1] * n
for u, v, i in w_edges:
if levels[u] == -1:
dfs(u, 0, -1)
for u, v, i in w_edges:
if i not in cri:
p_cri.add(i)
return [cri, p_cri]- Time complexity: O(ELogV), Where E is the no. of edges and V is no. of vertices.
- Space complexity: O(V) where V is no. of vertices.
Practice Question
Frequently Asked Questions
Q.1: What is the difference between Kruskal’s and Prim’s algorithms?
- Prims Algo: The algorithm obtains the minimum spanning tree by choosing the adjacent vertices from a set of selected vertices
- Kruskal Algo: To obtain the minimum spanning tree this algorithm selects the edges from a set of edges.
Q.2: How efficient is the Kruskal algorithm?
Ans: Kruskal performs better in typical situations (sparse graphs) because it uses simpler data structures and its time complexity is O(ELogV), Where E is the no. of edges and V is no. of vertices.
Q.3: Is Kruskal better than prim?
Ans: Prim’s algorithm is significantly faster in the limit when you’ve got a really dense graph with many more edges than vertices. Kruskal performs better in typical situations (sparse graphs) because it uses simpler data structures.